Principle components analysis
?princomp
data(iris)
# log transform
log.ir <- log(iris[, 1:4])
ir.species <- iris[, 5]
# apply PCA - scale. = TRUE is highly
# advisable, but default is FALSE.
ir.pca <- prcomp(log.ir,
center = TRUE,
scale. = TRUE) # plot method
plot(ir.pca, type = "l")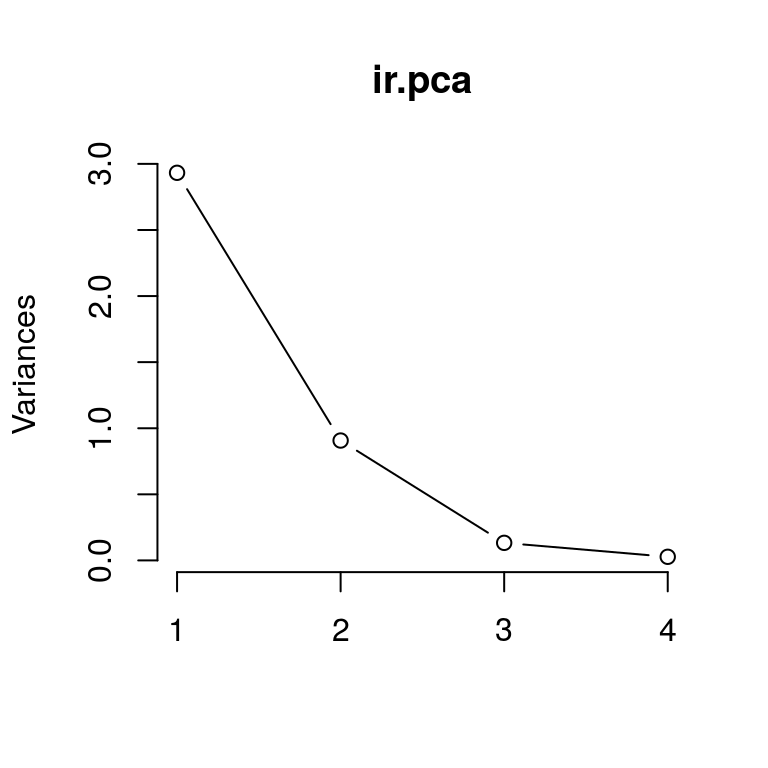
biplot(ir.pca)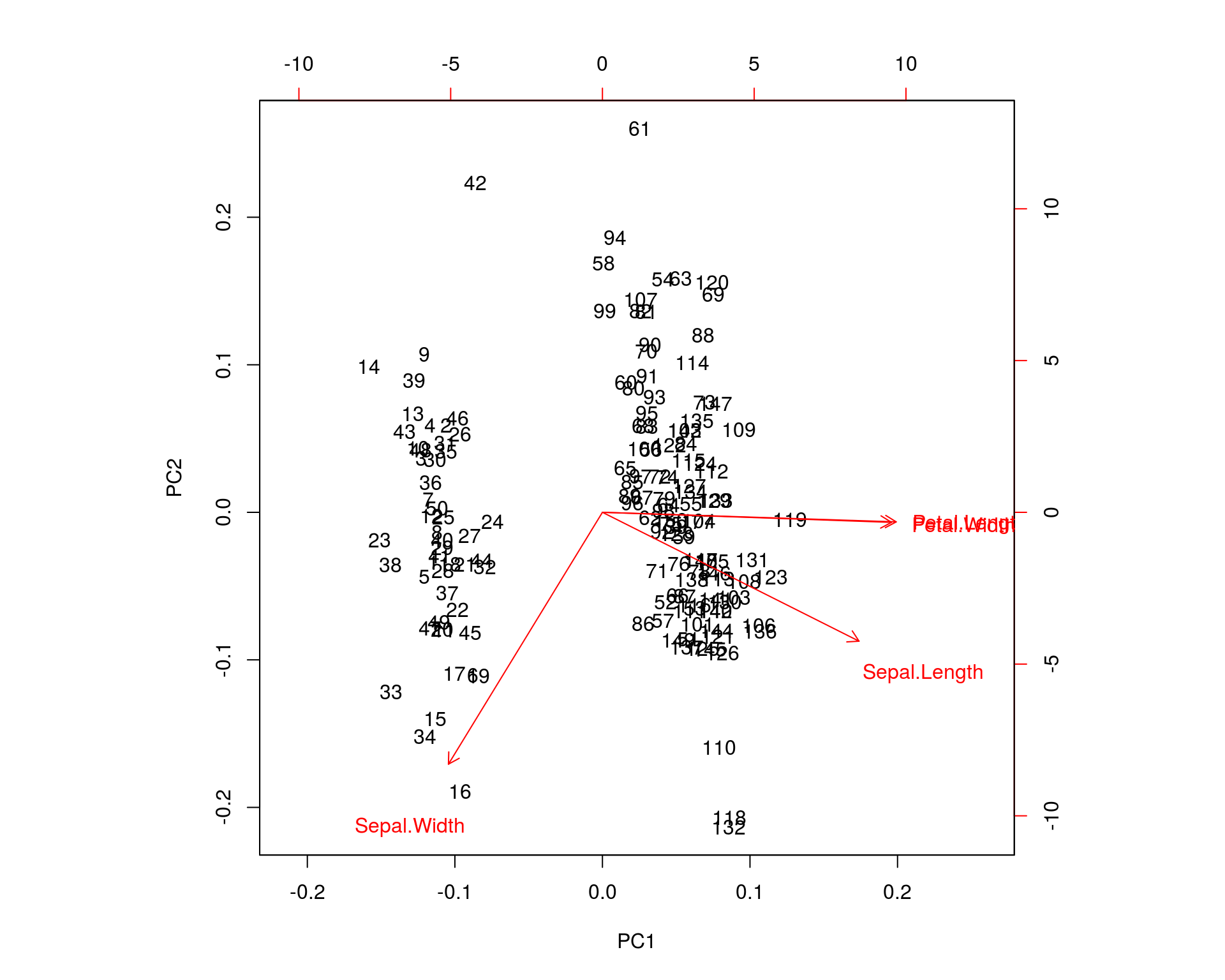
Links
https://www.r-bloggers.com/computing-and-visualizing-pca-in-r/
http://rcharlie.com/2017-04-13-Coachella/
Copyright © Brian J. Knaus. All rights reserved.